Solution Review
8 FREE Open Source SOFTWARE FOR IMAGE ALYSIS
Linux, OpenOffice, and GIMP are the first things that come to mind when I think about “Open Source Software”. But there are many other free programs available for different purposes, including image analysis (IA), of pathology slides.
This article describes the concept Open Source Software. It also discusses the 8 most widely used open-source image analysis software that can be usedalone or in combination for pathology to analyze whole slide images (WSI ).
The history of open source software is nearly as old as the history of software development. This method of creating software is based on sharing, collaborative improvement of source code . With the widespread recognition of Linux, an open-source operating system, and the release the Netscape browser’s source codes, the interest in open-source computer programs grew significantly. In 1998, the Open Source Initiative (OSI) was created.
What is “OPEN SOURCE SOFTWARE”?
Anyone can freely access, use, modify, and share open source software programs. Open source software can be made by many people, and distributed under licenses that conform to the Open Source Definition.
OPEN SOURCE LIENSES
Open source licenses conform with the Open Source Definition. They allow software to be freely modified and shared . A license must be reviewed by the Open Source Initiative before it can be approved by the Open Source Initiative.
You can find the list open-source license types here
To be considered “Open Source software”, any software license and software distributed under it must meet the following ten criteria:
- Redistribution at no cost
- The program must contain source code and allow distribution
- Modifications and derived works must be permitted
- Integrity of the Author’s Source Code
- No discrimination against persons or groups
- No discrimination against Fields of Endeavor
- Distribution of license
- A Product License must not be specific
- License must not restrict other software
- Technology-neutral Licenses are required
CAN OPEN SOFTWARE BE USEED FOR COMMERCIAL PURPOSE?
Yes, all Open Source software is allowed to be used commercially. The Open Source Definition allows for this. Open Source software can be sold.
commercial does NOT mean proprietary. You must distribute the software, or any modified version of it, under the same license as the original, i.e. You must make it open-source.
8 POPULAR OPEN SOURCE OPEN IMAGE ANALYSIS PROGRAMS THAT CAN BE USE IN PATHOLOGY
Image analysis algorithms are transforming life sciences by quantifying the visual information in pathology images. This list and description contains the 8 most widely used open-source programs for image analysis. They can both be used by novices and professionals alike. Each program is a standalone IA software that has its own strengths and weaknesses. However, many of them can be combined with one another via plugins (software components which can be installed separately to add functionality) to enhance their capabilities and address specific IA issues each project faces.
1. IMAGE J
ImageJ is a Java-based program for image processing. It was developed in collaboration between the National Institutes of Health NIH and Laboratory for Optical and Computational Instrumentation at the University of Wisconsin. ImageJ is the most well-known and longest-lived open-source software for biomedical imaging analysis. ImageJ is an experimental program and NIH is not responsible for any other users. The end user would need to validate and verify the program in order to use it in regulated environments.
A large number of ImageJ plugins has been created over the years, which covers most aspects of biological image analysis. While many of them are simple to use, some require programming skills.
ImageJ supports over 150 biological image formats, including WSI. The problem is with WSI. It’s not the format that is the issue, but the image size. A plugin called was created to solve this problem. This plugin is intended for rapid prototyping and testing algorithms on digital slides that are aimed at research.
TUTORIALS AND DOWNLOAD
The software website offers tutorials, and you can download the software here.
You will also find links to many presentations as well as key-note lectures on ImageJ. The user community has a large and active membership. You can also ask questions and receive feedback via the forum.
1A. FIJI (FIJI IS JUST IMAGE J)
Fiji is a Distribution of ImageJ. It’s a bundle of ImageJ plugins that is ready-to-use for life sciences. Fiji was established to help with the installation and maintenance TrakEM2, one of the more complicated ImageJ plugins. This provides complete solutions for managing, registration, segmentation and annotation large data sets from electron microscopy. Fiji was also home to other plugins that allow for advanced life science image analysis. These plugins, unlike ImageJ, are curated and organized into categories in the plugin menu. This makes them easier to find and use.
WSI PROCESSING CAPABILITIES
Yes with the SlideJ plug
LEVEL OFSKILLS:
This software is intended for researchers who have minimal computer skills. However, ImageJ functionality can easily be extended with plugins. It has also been appealing to researchers with software development skills .
HOW TO CITE
ImageJ and Fiji are funded based on citations. Please cite appropriate papers in the ImageJ Citation Guide.
TUTORIALS AND DOWNLOAD
Fiji can be downloaded from this link. Tutorials are available on the Fiji YouTube channel.
2. QUPATH
Designed at the Queen’s University Belfast QPath, is a free and open-source desktop software program specifically for analyzing WSI. Its primary purpose is IHC quantification (whole slides, tumor microarrays), but has also been used to analyze H&E tumors.
It also includes a user-friendly WSI reader with smart annotation tools that use pixel information in slides to speed up the process and increase precision (e.g. Semi-automatic delineation (based on contrast between epidermis & dermis) of the epidermis using extensive visualization tools.
QuPath provides ready-to use IA algorithms to solve common pathology problems, such as automatic cells classification for brightfield or fluorescent images. It also offers building blocks that can be linked together in order to create custom workflows. QuPath allows developers to create their own extensions and solve new problems.
QuPath’s functions include:
- Multi-slide projects created
- Stain estimation
- Cell detection and feature calculation
- Machine learning random trees classifier for trainable cell classification
- Batch processing
- Use survival analysis tools to link analysis results to clinical data to uncover the roles and biomarkers
- Fluorescence support
- Data exchange with existing platforms and software (e.g. ImageJ and MATLAB
- Scriptable data mining
- Rapid visualization, export, and generation of spatial, morphological, and intensity-based features
WSI PROCESSING CAPABILITIES –
Yes The program is designed to handle WSI
LEVEL OF SKILLS
This extension is for pathologists and researches without programming skills , but developers with software development skills can add their extensions.
HOW TO CITE
QuPath is open source software for digital pathology image analysis. Please cite Bankhead, P. and al. QuPath is an open source software program for digital pathology image analysis. Scientific Reports 2017 ).
TUTORIALS AND DOWNLOAD
QuPath is available for download here. The tutorials are also available on Pete Bankhead’s YouTube channel.
3. CELL PROFILER
Broad Institute of MIT, Harvard has developed cell profiler. This MATLAB-based open source software allows biologists and scientists to analyse and batch-process cells from biological images . It features a modular design as well as a user-friendly interface that allows users to point to complete most tasks. Each module can be combined into a pipeline, which then automatically analyzes the images. An common pipeline includes loading images, correcting uneven illumination, identifying objects and taking measurements. These modules can be easily added, removed or rearranged within a pipeline. Each module is executed in a specific order. One can see the performance of each module while they are being executed. It is possible to mix and match modules and adjust parameters according to the intended use. The analyze is object-based. It identifies the using criteria such as size and shape. Hierarchically, the objects are identified/generated – primary objects eg. Cell nuclei are used as a base to create secondary objects. For example, cells that contain the previously identified nuclei and their surrounding features. Measurements of features such as area, shape and color intensity, degree correlation between colors, texture (smoothness) and number of neighbors are made per object . The outline of the objects created are displayed over the original image to allow for quality control and verification. The program can extract metadata from images. A personal computer can process small numbers of images automatically, while a computing cluster can analyze hundreds of thousands of images. Although the software cannot be used for WSI analysis by itself, it can be integrated with other programs such as e.g. Orbit cuts the WSI into tiles, and sends them to Cell profiler for analysis.
WSI PROCESSING CAPABILITIES –
Integration with other programs (e.g. Orbit can only be integrated with other programs, e.g.
LEVEL OF SKILLS
This program is for researchers with no programming or computer vision background.
HOW TO CITE
Use Cell profiler to do your research. Please cite Anne E. Carpenter, Lamprecht, Michael R. and David M. Sabatini. Biotechniques 42.1 (2007), 71-75.
TUTORIALS AND DOWNLOAD
The Cell profiler software can be downloaded from this link.
3A. 3A.
CellProfiler Analyst software is designed to explore data generated by CellProfiler. This software data analysis software is not for image analysis.
TUTORIALS AND DOWNLOAD
Cell profiler analyst can downloaded here, and the tutorials can be found here.
HOW TO CITE
CellProfiler Analysts is data exploration and analysis software that can be used to analyze complex images. BMC Bioinformatics 9(1).482
4. ILASTIK
Ilastik, an simple open-source tool that allows users to segment and classify 2, 3, and 4D images in a consistent way is Ilastik. Using a random forests classifier, ilastik uses labels supplied by users to learn from them. Using these labels, ilastik determines the problem and applies a segmentation.
Ilastik can support up to three spatial and one spectral dimensions. It makes full use of all dimensions for feature calculation. The program provides real time feedback which allows the user to interactively define the segmentation results.
The program can improve classification based upon user feedback. The classifier will make a prediction that may indicate where it might be wrong. The prediction is displayed on an overlay called uncertaintymap, which highlights the regions where the prediction is most unclear. It helps the user find the most difficult areas of an image, and where the classifier considers the new labels to be most informative. After ilastik is trained on a sample set of images, it can automatically process large numbers of images using either the batch processing mode of CellProfiler or the pipeline . A plug-in mechanism allows you to add new analysis modules.
Future versions will include:
- Seeded segmentation algorithms, such as watershed
- Image features and multi-spectral images can be condensed using unsupervised dimension reduction techniques
- Hierarchical processing allows for subsequent analyses on top intermediate results
Ilastik can only capture local cues like color, brightness, and texture . is not intended to capture global configurations. Therefore, analysis of spatial relationships between objects will not possible.
The program is not intended to handle entire slide images, but it has been used in pathology to detect mitotic characters in high power fields.
WSI PROCESSING CAPABILITIES
It is not but can be used for high-power fields analysis of pathology slides.
LEVEL OF SKILLS
This is for researchers who don’t have any computer programming skills. However, an understanding of image processing will be helpful.
HOW TO CITE
When using ilastik for your analysis cite: Sommer, Christoph, et al. “Ilastik – Interactive learning and segmentation toolkit.” 2011 IEEE international symposium on Biomedical Imaging: From nano to micro. IEEE, 2011.
TUTORIALS AND DOWNLOAD
You can download the software here. The tutorials are also available on the ilastik team YouTube channel. A forum is also available to users. Here they can discuss and ask questions.
5. ORBIT
Orbit Image Analysis, a free open-source software that quantifies large-format images like whole slides of tissue, can connect to Open Microscopy Environment (Omero) or load them from local disk. Orbit Image Analysis can also process images on local computers or clusters using the Spark job server.
Orbit can be used for defining a region or interest (ROI), in WSI. A tile-based process pipeline makes it possible to process WSI. It also allows for different resolutions of the image. To process individual cells within each tile, sub-regions of tiled can be sent later to CellProfiler. CellProfiler processes all tiles within the region of interest. The results are returned into Orbit, where they can be visualized.
The software includes many embedded image analysis algorithms for tissue quantify using machine-learning techniques, cell segmentation and object classification, to name just a few.
Object detection is based on trainable foreground/background classes. Overlapping objects (e.g. Cell clusters can be broken down afterward. Features of objects strong>Features of objects (e.g.
Segmented objects may be classified based on their features such as size and shape factors, or intensities . It can, for example, distinguish between unstained and stained cells and between round and edgy items. This classification uses machine learning. The user can select examples to specify classes.
ROIs are defined either by hand annotations or using a trainable exclusionmap that defines the “good” areas where analysis should take place. Both methods of defining ROIs are possible to combine.
Orbit has ready-to-use models for lung fibrosis in rats and glomeruli detection and treatment in mice and rats.
Users who have software development skills are able to create scripts and then apply them all tiles in the ROI. This program can also be used with ImageJ, and its plugins for analysis.
THE INTEGRATION of ORBIT WITH CELLPROFILER. LET US USE CELLPROFILER’S IMAGE ANALYSIS FUNCTIONALITIES ON THE TILED WSIS.
Orbit can be used with any format from the bioformats library, even though it is optimized for WSI formats (such as: SVS and NDPI, NDPIS SCN, AFI. CZI. IMSI, VSI. ETSI.
WSI PROCESSING CAPABILITIES
Yes
LEVEL OF SKILLS
This extension is for researchers who don’t have any computer skills, but developers with programming skills can create their own extensions.
HOW TO CITE
I didn’t find any official recommendations for which publication I should cite. However, if I were using it for research, I would cite the following: Goldberg, Ilya, and others. “The Open Microscopy Environment’s (OME) Data Model, and XML File: Open Tools for Informatics and Quantitative Analysis in Biological Imaging” Genome Biology 6.5 (2006): R47.
TUTORIALS AND DOWNLOAD
The software can be downloaded here: https://www.orbit.bio/ and tutorial videos and different presentations about OME can be found on the OME YouTube channel https://www.youtube.com/channel/UCyySB9ZzNi8aBGYqcxSrauQ/videos
6. CYTOMINE
Cytomine, an open-source web-based image analysis and management tool is Cytomine. It was developed for active collaboration between multidisciplinary teams to large-scale image-based research.
The program supports different microscopy formats ( , including most of the whole slide scanner formats), either directly accessing the native format or converting to a pyramidal format during upload phase.
Cytomine allows to organize, explore and share (semantically as well as quantitatively) multigigapixel (including whole slide pathology images ) imaging data via the internet.
This program is ideal for collaborative projects and allows for permission management at both system and project levels.
Images can be uploaded from the user’s computer to Cytomine and stored in the cloud. They can later be organized in projects and explored in a Web-based viewer. The images can then be annotated and shared with others, reviewed and analyzed using both standard and custom algorithms.
Software and viewer features that are interesting include:
- Multiple images can be viewed simultaneously
- Multiple collaborators can access slides and annotations simultaneously
- Annotation layers are separate for each contributor
- Ability to search and sort annotations using different attributes and filters (including those generated automatically by an algorithm and not manually).
- Annotations for deep learning model development
- Conversion of model and algorithm results into editable annotations which can later be used to develop a new model
- Based on the resolution of the image, calculate area in mm
- Keywords (tags, terms and properties) are used to search for projects
- Create “image groups” that contain different stains from serial sections of the same block
WSI capabilities
Yes.
Level of skill
This tool is for researchers who don’t have computer skills (annotations, included deployable algorithms, and models) , developers with software development skills who can create their own algorithms (scripts and apps in Python, Java, JavaScript, or import algorithm from other software platforms, e.g. ImageJ, CellProfiler)
How do I cite?
Paper: Collaboration analysis of multi-gigapixel imagery data using Cytomine
Tutorials and Access
The program can be accessed here: https://doc.cytomine.org/user-guide/getting-started#get-an-account
The tutorials are available on the Neubias YouTube Channel.
Part 2: focusing on image analysis and data management
7. ICY
France-BioImaging and Institut Pasteur created Icy, a free open source software. It is a “collaborative photoshop dedicated to image analyses”, according to its website. Collaboration means that users who have software development skills may add features to the program. Icy allows users to visualize, annotate, and quantify bioimaging data. It was created as a common platform for image analysts who can create new algorithms or life scientists searching for an intuitive tool to image analysis.
The GUI was designed using components that users are already familiar with. For example, the main ‘ribbon toolbar looks similar to Office Suite (Microsoft) and provides access all functions of Icy.
LEVEL OF SKILLS
This tool is intended for software developers but can be used by anyone without programming skills.
HOW TO CITE
Cite De Chaumont, Fabrice, and co-authors when using Icy in your research. Nature methods 9.7 (2012), 690.
TUTORIALS AND DOWNLOAD:
The program can be downloaded here and the tutorials can also be found on Ice’s YouTube channel
8. PATHML
PathML isn’t a traditional image analysis program. It is a free, open-source toolkit for creating images in different formats and using different imaging modalities. It was created by researchers from the Dana-Farber Cancer Institute, Boston, MA, USA.
PathML is a framework that allows you to create and run preprocessing pipelines for different file formats.
Python 3 is used to write the PathML library.
PathML supports several imaging modalities (H&E, IHC, IF, 3-D images etc. ) There are 160+ popular image formats, including tiff and OME-TIFF.
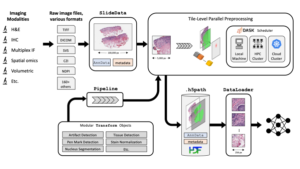 The first step in a PathML workflow involves loading the raw image file and creating a SlideData object. This is the central data type that represents an image with associated metadata.
The first step in a PathML workflow involves loading the raw image file and creating a SlideData object. This is the central data type that represents an image with associated metadata.
Next, create a preprocessing pipeline. This is the sequential application and transformation of independent building blocks or image tiles. The images can then be applied using the independently developed image algorithms.
WSI capabilities
Yes.
Level of skill
This is for developers with software development skills.
How do I cite
Code and documentation
The code can be found on Github: https://github.com/Dana-Farber-AIOS/pathml and the documentation and example workflows can be found on the PathML project page: https://pathml.org/
SUMMARY
Image analysis of pathology specimens is a rapidly growing area of biomedical research. This niche is a popular one, with many companies offering high-value software or services. Image analysis doesn’t have to be costly. Many open-source programs with advanced image analysis capabilities are available many of them are specifically developed to handle WSI.
These are the eight most easily accessible images that can be used to analyze pathology images
- ImageJ/ Fiji
- QuPath
- CellProfiler
- Ilastik
- Orbit
- Cytomine
- Icy
- PathML
You can find more information on different image analysis programs at the Bioimage Information Search Engine (BISE). This site, which is similar to Wikipedia for image analysis, contains curated information about numerous commercial and open-source image analysis programs.
REFERENCES
- Schneider, Caroline A. Wayne S. Rasband and Kevin W. Eliceiri. “NIH ImageJ: 25 Years of Image Analysis.” Nature Methods7 (2012): 671.
- Rueden, Curtis T., et al. “ImageJ2 – ImageJ for next generation scientific image data.” BMC Bioinformatics1 (2018): 529.
- Bankhead, Peter, et al. “QuPath” Scientific Reports1 (2017): 16788
- Lamprecht, Michael R. and Anne E. Carpenter. “CellProfiler(tm), a free and versatile software for automated bioimage analysis.” Biotechniques1 (2007) 71-75.
- Jones, Thouis R., et al. “CellProfiler Analyzer: Data Exploration and Analysis Software for Complex Image-Based Screens.” BMC Bioinformatics1 (2008) 482.
- Sommer, Christoph, et al. “Ilastik”: Interactive learning and segmentation software. 2011 IEEE International Symposium on Biomedical Imaging: From nano to micro. IEEE, 2011.
- Goldberg, Ilya G., et al. “The Open Microscopy Environment, (OME), Data Model and XML File: Open Tools for Informatics and Quantitative Analysis in Biological Imaging.” Genome Biology5 (2005). R47.
- Maree, Raphael, et al. Maree, Raphael, & Co.
- De Chaumont, Fabrice, et al. “Icy: An open bioimage informatics platform to extend reproducible research.” 7 (2012): 690.
- Rosenthal, Jacob, Ryan Carelli, Mohamed Omar, David Brundage, Ella Halbert, Jackson Nyman, Surya N. Hari et al. “Building tools to machine learning and artificial Intelligence in Cancer Research: Best Practices and a Case Study with the PathML Toolkit for Computational Pathology” Molecular Cancer Research 20. 2 (2022): 202-206.










